Origin as a Primitive Prokaryotic Immune System:
Prokaryotic organisms are in a continual evolutionary arms-race with bacteriophages, which may parasitize a prokaryote by injecting its own viral DNA/RNA and using its host cell’s machinery to replicate itself. One adaptation that many bacterial species have developed to combat viral challenges is the CRISPR/cas9 system. CRSIPR (clustered regularly interspersed short palindromic repeats) sequences have been observed in the bacterial genome and contain “spacer” sequences that are derived from the genetic material injected by viruses. This allows a prokaryote to hold a primitive acquired immune system against the viruses present in its environment. With the injection of DNA/RNA, a prokaryote may incorporate the phage’s genetic material into its own genome, becoming one of the spacers within the CRISPR sequence. Common within CRISPR sequences is also a gene coding for a cas (CRISPR-associated) protein. The phage-specific spacer sequence is then transcribed to mRNA, and the cas9 gene is transcribed and translated to produce the functional protein. Using the phage-specific mRNA as a template, the cas9 protein uses this template to identify viral DNA and cleave it to produce a double-strand break. In a cell’s effort to repair a double-strand break (nonhomologous end-joining), frameshift mutations typically arise, rendering the viral DNA inactive and unable of creating further viruses. In this way, the bacterium has acquired an immunity to that specific phage through the CRIPSR/cas9 system (Barrangou et al. 2007).
Re-purposing the CRISPR/cas9 System in Higher Organisms:
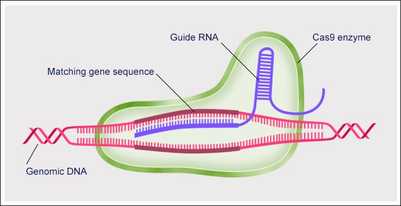
In recent years, the CRISPR/cas9 system has emerged as an efficient method of gene modification and gene knock-out in eukaryotes. To study the role of a specific gene, one must synthesize a “guide RNA” to take the place of the phage-specific spacer sequence in prokaryotes. A guide RNA is composed of three elements: a target sequence (specific to the gene you wish to study), a PAM (protospacer adjacent motif) element, and an identification element (general to all guide RNAs, used for identification of the RNA by the cas9 protein). The PAM element (5’-NGG-3’) is critical for targeting viral DNA as opposed to genomic DNA, and its absence will lead to an unsuccessful cleavage. The specific PAM sequence, however, is located within the target gene that you wish to knock-down, and thus the guide RNA needs to contain a sequence complimentary to the PAM sequence, bridging the target and identification elements. After successful synthesis of the guide RNA, its introduction along with the cas9 protein will produce a double-strand break in your target gene, causing a frameshift mutation and knocking-out expression.
References:
http://www.abc.net.au/cm/lb/7238280/data/crispr-cas9-at-work-data.jpg
Prokaryotic organisms are in a continual evolutionary arms-race with bacteriophages, which may parasitize a prokaryote by injecting its own viral DNA/RNA and using its host cell’s machinery to replicate itself. One adaptation that many bacterial species have developed to combat viral challenges is the CRISPR/cas9 system. CRSIPR (clustered regularly interspersed short palindromic repeats) sequences have been observed in the bacterial genome and contain “spacer” sequences that are derived from the genetic material injected by viruses. This allows a prokaryote to hold a primitive acquired immune system against the viruses present in its environment. With the injection of DNA/RNA, a prokaryote may incorporate the phage’s genetic material into its own genome, becoming one of the spacers within the CRISPR sequence. Common within CRISPR sequences is also a gene coding for a cas (CRISPR-associated) protein. The phage-specific spacer sequence is then transcribed to mRNA, and the cas9 gene is transcribed and translated to produce the functional protein. Using the phage-specific mRNA as a template, the cas9 protein uses this template to identify viral DNA and cleave it to produce a double-strand break. In a cell’s effort to repair a double-strand break (nonhomologous end-joining), frameshift mutations typically arise, rendering the viral DNA inactive and unable of creating further viruses. In this way, the bacterium has acquired an immunity to that specific phage through the CRIPSR/cas9 system (Barrangou et al. 2007).
Re-purposing the CRISPR/cas9 System in Higher Organisms:
In recent years, the CRISPR/cas9 system has emerged as an efficient method of gene modification and gene knock-out in eukaryotes. To study the role of a specific gene, one must synthesize a “guide RNA” to take the place of the phage-specific spacer sequence in prokaryotes. A guide RNA is composed of three elements: a target sequence (specific to the gene you wish to study), a PAM (protospacer adjacent motif) element, and an identification element (general to all guide RNAs, used for identification of the RNA by the cas9 protein). The PAM element (5’-NGG-3’) is critical for targeting viral DNA as opposed to genomic DNA, and its absence will lead to an unsuccessful cleavage. The specific PAM sequence, however, is located within the target gene that you wish to knock-down, and thus the guide RNA needs to contain a sequence complimentary to the PAM sequence, bridging the target and identification elements. After successful synthesis of the guide RNA, its introduction along with the cas9 protein will produce a double-strand break in your target gene, causing a frameshift mutation and knocking-out expression.
References:
- Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P. 2007. CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes. Science 315: 1709-1712.
http://www.abc.net.au/cm/lb/7238280/data/crispr-cas9-at-work-data.jpg
 RSS Feed
RSS Feed
